Code examples¶
Applications¶
Creep compliance analysis¶
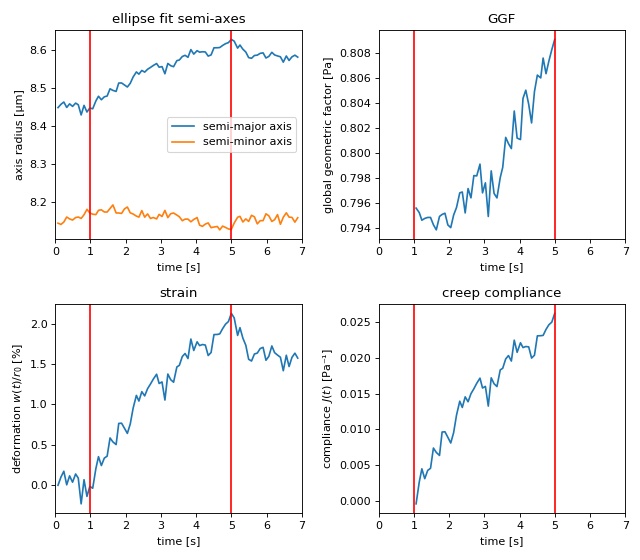
This example uses the contour data of an HL60 cell in the OS to compute its GGF and creep compliance. The contour data were determined from this phase-contrast video (prior to video compression). During stretching, the total laser power was increased from 0.2W to 1.3W (reflexes due to second harmonic effects appear as white spots).
1import ggf
2import h5py
3import lmfit
4import matplotlib.pylab as plt
5import numpy as np
6
7
8def ellipse_fit(radius, theta):
9 '''Fit a centered ellipse to data in polar coordinates
10
11 Parameters
12 ----------
13 radius: 1d ndarray
14 radial coordinates
15 theta: 1d ndarray
16 angular coordinates [rad]
17
18 Returns
19 -------
20 a, b: floats
21 semi-axes of the ellipse; `a` is aligned with theta=0.
22 '''
23 def residuals(params, radius, theta):
24 a = params["a"].value
25 b = params["b"].value
26 r = a*b / np.sqrt(a**2 * np.sin(theta)**2 + b**2 * np.cos(theta)**2)
27 return r - radius
28
29 parms = lmfit.Parameters()
30 parms.add(name="a", value=radius.mean())
31 parms.add(name="b", value=radius.mean())
32
33 res = lmfit.minimize(residuals, parms, args=(radius, theta))
34
35 return res.params["a"].value, res.params["b"].value
36
37
38# load the contour data (stored in polar coordinates)
39with h5py.File("data/creep_compliance_data.h5", "r") as h5:
40 radius = h5["radius"][:] * 1e-6 # [µm] to [m]
41 theta = h5["theta"][:]
42 time = h5["time"][:]
43 meta = dict(h5.attrs)
44
45
46factors = np.zeros(len(radius), dtype=float)
47semimaj = np.zeros(len(radius), dtype=float)
48semimin = np.zeros(len(radius), dtype=float)
49strains = np.zeros(len(radius), dtype=float)
50complnc = np.zeros(len(radius), dtype=float)
51
52for ii in range(len(radius)):
53 # determine semi-major and semi-minor axes
54 smaj, smin = ellipse_fit(radius[ii], theta[ii])
55 semimaj[ii] = smaj
56 semimin[ii] = smin
57 # compute GGF
58 if (time[ii] > meta["time_stretch_begin [s]"]
59 and time[ii] < meta["time_stretch_end [s]"]):
60 power_per_fiber = meta["power_per_fiber_stretch [W]"]
61 f = ggf.get_ggf(
62 model="boyde2009",
63 semi_major=smaj,
64 semi_minor=smin,
65 object_index=meta["object_index"],
66 medium_index=meta["medium_index"],
67 effective_fiber_distance=meta["effective_fiber_distance [m]"],
68 mode_field_diameter=meta["mode_field_diameter [m]"],
69 power_per_fiber=power_per_fiber,
70 wavelength=meta["wavelength [m]"],
71 poisson_ratio=.5)
72 else:
73 power_per_fiber = meta["power_per_fiber_trap [W]"]
74 f = np.nan
75 factors[ii] = f
76
77# compute compliance
78strains = (semimaj-semimaj[0]) / semimaj[0]
79complnc = strains / factors
80compl_ival = (time > meta["time_stretch_begin [s]"]) * \
81 (time < meta["time_stretch_end [s]"])
82stretch_index = np.where(compl_ival)[0][0]
83complnc_1 = strains/factors[stretch_index]
84
85# plots
86plt.figure(figsize=(8, 7))
87
88ax1 = plt.subplot(221, title="ellipse fit semi-axes")
89ax1.plot(time, semimaj*1e6, label="semi-major axis")
90ax1.plot(time, semimin*1e6, label="semi-minor axis")
91ax1.legend()
92ax1.set_xlabel("time [s]")
93ax1.set_ylabel("axis radius [µm]")
94
95ax2 = plt.subplot(222, title="GGF")
96ax2.plot(time, factors)
97ax2.set_xlabel("time [s]")
98ax2.set_ylabel("global geometric factor [Pa]")
99
100ax3 = plt.subplot(223, title="strain")
101ax3.plot(time, (strains)*100)
102ax3.set_xlabel("time [s]")
103ax3.set_ylabel("deformation $w(t)/r_0$ [%]")
104
105ax4 = plt.subplot(224, title="creep compliance")
106ax4.plot(time[compl_ival], complnc[compl_ival])
107ax4.set_xlabel("time [s]")
108ax4.set_ylabel("compliance $J(t)$ [Pa⁻¹]")
109
110for ax in [ax1, ax2, ax3, ax4]:
111 ax.set_xlim(0, np.round(time.max()))
112 ax.axvline(x=meta["time_stretch_begin [s]"], c="r")
113 ax.axvline(x=meta["time_stretch_end [s]"], c="r")
114
115plt.tight_layout()
116plt.show()
Reproduction tests¶
Radial stresses of a prolate spheroid¶
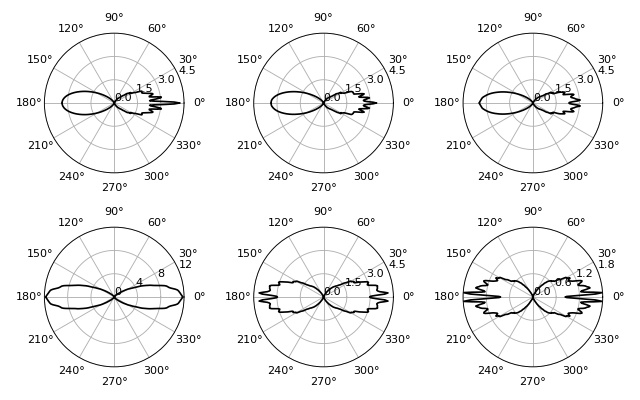
This examples computes radial stress profiles for spheroidal objects in the optical stretcher, reproducing figures (9) and (10) of [BCG09].
1import matplotlib.pylab as plt
2import numpy as np
3import percache
4
5from ggf.stress.boyde2009.core import stress
6
7
8@percache.Cache("stress_reproduced.cache", livesync=True)
9def compute(**kwargs):
10 "Locally cached version of ggf.core.stress"
11 return stress(**kwargs)
12
13
14# variables from the publication
15alpha = 47
16wavelength = 1064e-9
17radius = alpha * wavelength / (2 * np.pi)
18
19kwargs = {"stretch_ratio": .1,
20 "object_index": 1.375,
21 "medium_index": 1.335,
22 "wavelength": wavelength,
23 "beam_waist": 3,
24 "semi_minor": radius,
25 "power_left": 1,
26 "power_right": 1,
27 "poisson_ratio": 0,
28 "n_points": 200,
29 }
30
31kwargs1 = kwargs.copy()
32kwargs1["power_right"] = 0
33kwargs1["stretch_ratio"] = 0
34kwargs1["dist"] = 90e-6
35
36kwargs2 = kwargs.copy()
37kwargs2["power_right"] = 0
38kwargs2["stretch_ratio"] = .05
39kwargs2["dist"] = 90e-6
40
41kwargs3 = kwargs.copy()
42kwargs3["power_right"] = 0
43kwargs3["stretch_ratio"] = .1
44kwargs3["dist"] = 90e-6
45
46kwargs4 = kwargs.copy()
47kwargs4["dist"] = 60e-6
48
49kwargs5 = kwargs.copy()
50kwargs5["dist"] = 120e-6
51
52kwargs6 = kwargs.copy()
53kwargs6["dist"] = 200e-6
54
55
56# polar plots
57plt.figure(figsize=(8, 5))
58
59th1, sigma1 = compute(**kwargs1)
60ax1 = plt.subplot(231, projection='polar')
61ax1.plot(th1, sigma1, "k")
62ax1.plot(th1 + np.pi, sigma1[::-1], "k")
63
64th2, sigma2 = compute(**kwargs2)
65ax2 = plt.subplot(232, projection='polar')
66ax2.plot(th2, sigma2, "k")
67ax2.plot(th2 + np.pi, sigma2[::-1], "k")
68
69th3, sigma3 = compute(**kwargs3)
70ax3 = plt.subplot(233, projection='polar')
71ax3.plot(th3, sigma3, "k")
72ax3.plot(th3 + np.pi, sigma3[::-1], "k")
73
74for ax in [ax1, ax2, ax3]:
75 ax.set_rticks([0, 1.5, 3, 4.5])
76 ax.set_rlim(0, 4.5)
77
78th4, sigma4 = compute(**kwargs4)
79ax4 = plt.subplot(234, projection='polar')
80ax4.plot(th4, sigma4, "k")
81ax4.plot(th4 + np.pi, sigma4[::-1], "k")
82ax4.set_rticks([0, 4, 8, 12])
83ax4.set_rlim(0, 12)
84
85th5, sigma5 = compute(**kwargs5)
86ax5 = plt.subplot(235, projection='polar')
87ax5.plot(th5, sigma5, "k")
88ax5.plot(th5 + np.pi, sigma5[::-1], "k")
89ax5.set_rticks([0, 1.5, 3, 4.5])
90ax5.set_rlim(0, 4.5)
91
92th6, sigma6 = compute(**kwargs6)
93ax6 = plt.subplot(236, projection='polar')
94ax6.plot(th6, sigma6, "k")
95ax6.plot(th6 + np.pi, sigma6[::-1], "k")
96ax6.set_rticks([0, 0.6, 1.2, 1.8])
97ax6.set_rlim(0, 1.8)
98
99for ax in [ax1, ax2, ax3, ax4, ax5, ax6]:
100 ax.set_thetagrids(np.linspace(0, 360, 12, endpoint=False))
101
102plt.tight_layout()
103plt.show()
Decomposition of stress in Legendre polynomials¶
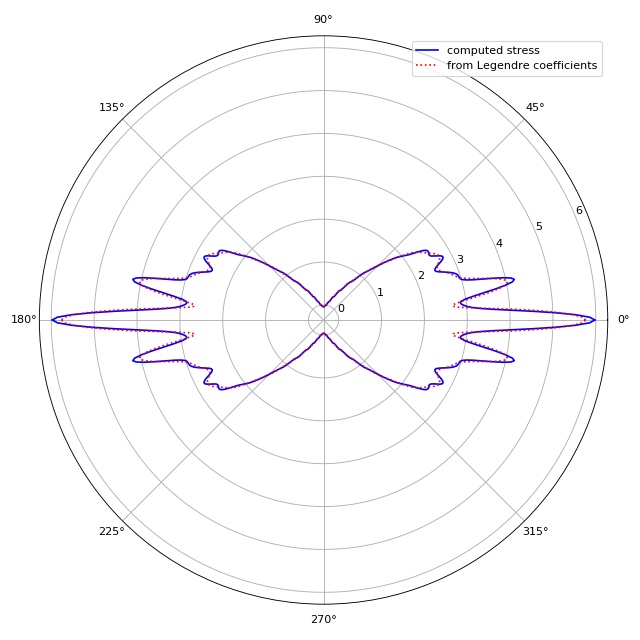
To compute the GGF, ggf.globgeomfact.coeff2ggf() uses the
coefficients of the decomposition of the stress into Legendre
polynomials \(P_n(\text{cos}(\theta))\). This example visualizes
the small differences between the original stress and the stress
computed from the Legendre coefficients. This plot is automatically
produced by the original Matlab script StretcherNStress.m.
Note that the original Matlab yields different results for the
same set of parameters, because the Poisson’s ratio (keyword
argument poisson_ratio) is non-zero;
see issue #1.
1import matplotlib.pylab as plt
2import numpy as np
3import percache
4
5from ggf.sci_funcs import legendrePlm
6from ggf.stress.boyde2009.core import stress
7
8
9@percache.Cache("stress_decomposition.cache", livesync=True)
10def compute(**kwargs):
11 "Locally cached version of ggf.core.stress"
12 return stress(**kwargs)
13
14
15# compute default stress
16theta, sigmarr, coeff = compute(ret_legendre_decomp=True,
17 n_points=300)
18
19# compute stress from coefficients
20numpoints = theta.size
21sigmarr_c = np.zeros((numpoints, 1), dtype=float)
22for ii in range(numpoints):
23 for jj, cc in enumerate(coeff):
24 sigmarr_c[ii] += coeff[jj] * \
25 np.real_if_close(legendrePlm(0, jj, np.cos(theta[ii])))
26
27# polar plot
28plt.figure(figsize=(8, 8))
29ax = plt.subplot(111, projection="polar")
30plt.plot(theta, sigmarr, '-b', label="computed stress")
31plt.plot(theta + np.pi, sigmarr[::-1], '-b')
32plt.plot(theta, sigmarr_c, ':r', label="from Legendre coefficients")
33plt.plot(theta + np.pi, sigmarr_c[::-1], ':r')
34plt.legend()
35
36plt.tight_layout()
37plt.show()
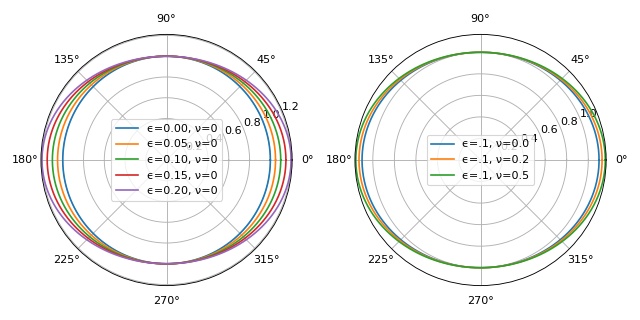
Object boundary: stretching and Poisson’s ratio¶
This example illustrates how the parameters Poisson’s ratio
\(\nu\) and stretch ratio \(\epsilon\) influence
the object boundary used in ggf.core.stress() and
defined in ggf.core.boundary().
Note that the boundary function was not defined correctly prior to version 0.3.0 (issue #1). Since version 0.3.0, the semi-minor axis is equivalent to the keyword argument a (1 by default).
1import numpy as np
2import matplotlib.pylab as plt
3
4from ggf.stress.boyde2009.core import boundary
5
6theta = np.linspace(0, 2*np.pi, 300)
7costheta = np.cos(theta)
8
9# change epsilon
10eps = [.0, .05, .10, .15, .20]
11b1s = []
12for ep in eps:
13 b1s.append(boundary(costheta=costheta,
14 epsilon=ep,
15 nu=.0))
16
17# change Poisson's ratio
18nus = [.0, .25, .5]
19b2s = []
20for nu in nus:
21 b2s.append(boundary(costheta=costheta,
22 epsilon=.1,
23 nu=nu))
24
25# plot
26plt.figure(figsize=(8, 4))
27
28ax1 = plt.subplot(121, projection="polar")
29for ep, bi in zip(eps, b1s):
30 ax1.plot(theta, bi, label="ϵ={:.2f}, ν=0".format(ep))
31ax1.legend()
32
33ax2 = plt.subplot(122, projection="polar")
34for nu, bi in zip(nus, b2s):
35 ax2.plot(theta, bi, label="ϵ=.1, ν={:.1f}".format(nu))
36ax2.legend()
37
38plt.tight_layout()
39plt.show()